Ligand-X Demo
Demo: Absolute Binding Free Energy (ABFE)
Video Walkthrough
Coming Soon: Video walkthrough demonstrating the complete ABFE workflow in Ligand-X.
Data Flow Pipeline
When you submit an ABFE calculation, your data flows through these stages:
1. Ligand Preparation
- Parse SDF/SMILES, add hydrogens
- Assign AM1-BCC partial charges
- Create OpenFE SmallMoleculeComponent
2. Protein Preparation
- Parse PDB, clean structure
- Create OpenFE ProteinComponent
3. Protocol Setup
- Configure AbsoluteBindingProtocol
- Set lambda windows (typically 11–20)
- Configure HREX parameters
4. Complex Leg
- Ligand bound to protein
- Gradually "turn off" ligand interactions
- Run HREX across lambda windows
- Compute ΔG_complex
5. Solvent Leg
- Ligand in water box
- Gradually "turn off" ligand interactions
- Run HREX across lambda windows
- Compute ΔG_solvent
6. Thermodynamic Cycle
- ΔG_binding = ΔG_complex − ΔG_solvent
- MBAR analysis for optimal estimate
- Error propagation for uncertainty
Results
- ΔG binding free energy (kcal/mol)
- Statistical uncertainty
- Overlap matrices showing sampling quality
- Convergence plots over simulation time
- Replica exchange statistics
The Two-Leg Calculation
ABFE uses a thermodynamic cycle with two parallel simulations:
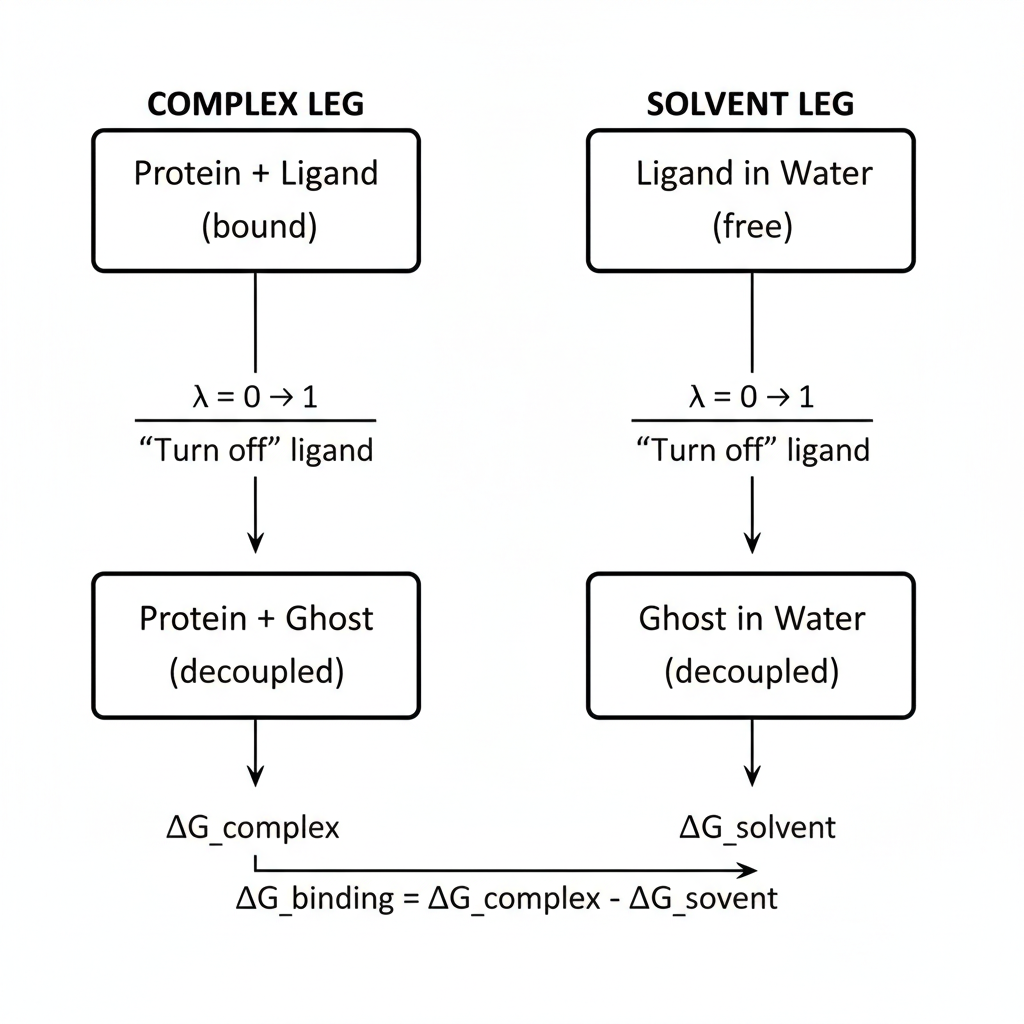
Figure 1: Thermodynamic cycle for ABFE calculations
Tools Used
| Stage | Tool | Purpose |
|---|---|---|
| Ligand Prep | RDKit | Molecule parsing, 3D generation |
| Charges | AM1-BCC | Partial charge assignment |
| Protein Prep | PDBFixer | Structure cleaning |
| Simulation | OpenMM | GPU-accelerated MD |
| Free Energy | OpenFE | AbsoluteBindingProtocol |
| Sampling | HREX | Hamiltonian Replica Exchange |
| Analysis | MBAR | Multistate Bennett Acceptance Ratio |
Example Output
After calculation completes, you’ll see:
- Binding Free Energy: ΔG in kcal/mol with error estimate
- Overlap Matrices: Heatmaps showing sampling quality between λ windows
- Convergence Plot: How the ΔG estimate stabilized over time
- Replica Exchange Stats: Acceptance rates for configuration swaps
- Thermodynamic Breakdown: Individual leg contributions
Protocol Settings
| Parameter | Default | Description |
|---|---|---|
simulation_time_ns |
5.0 | Production simulation per replica |
n_lambda_windows |
11 | Number of alchemical states |
equilibration_length_ns |
1.0 | Equilibration time per replica |
n_replicas |
11 | Number of HREX replicas |