projects – Full Stack
 Ligand-X is a web application for structural, physical, and chemical analysis of ligand-protein complexes, lowering the barrier to entry for advanced bioinformatics and cheminformatics simulations.
Read More ›
Ligand-X is a web application for structural, physical, and chemical analysis of ligand-protein complexes, lowering the barrier to entry for advanced bioinformatics and cheminformatics simulations.
Read More ›
blog – Binding Free Energy
 RBFE calculations let us compare how strongly different molecules bind to a protein target, helping medicinal chemists prioritize which compounds to synthesize next.
Read More ›
RBFE calculations let us compare how strongly different molecules bind to a protein target, helping medicinal chemists prioritize which compounds to synthesize next.
Read More ›
blog – Binding Free Energy
 ABFE calculations determine the exact binding strength of a drug molecule to its protein target—the gold standard measurement for understanding how potent a compound really is.
Read More ›
ABFE calculations determine the exact binding strength of a drug molecule to its protein target—the gold standard measurement for understanding how potent a compound really is.
Read More ›
blog – Biophysics simulation
 Molecular Dynamics (MD) simulations allow us to study the physical movements of atoms and molecules, providing a view of the dynamic evolution of the system.
Read More ›
Molecular Dynamics (MD) simulations allow us to study the physical movements of atoms and molecules, providing a view of the dynamic evolution of the system.
Read More ›
blog – Computational Drug Design
 Drug-like chemical space contains an estimated $10^{60}$ molecules, which is far more than can ever be synthesised and tested feasibly. Molecular docking provides a quick in silico filter, screening millions of candidates in hours and returning a ranked shortlist that guides what chemists synthesise.
Read More ›
Drug-like chemical space contains an estimated $10^{60}$ molecules, which is far more than can ever be synthesised and tested feasibly. Molecular docking provides a quick in silico filter, screening millions of candidates in hours and returning a ranked shortlist that guides what chemists synthesise.
Read More ›
blog – Garbage In, Garbage Out
 Before any protein simulation can begin, the raw PDB structure must be cleaned and prepared. This involves removing water molecules, adding missing hydrogens, and fixing incomplete residues. Similarly, small molecule ligands must be converted from SMILES / 3D coordinates to a chemically accurate 3D model.
Read More ›
Before any protein simulation can begin, the raw PDB structure must be cleaned and prepared. This involves removing water molecules, adding missing hydrogens, and fixing incomplete residues. Similarly, small molecule ligands must be converted from SMILES / 3D coordinates to a chemically accurate 3D model.
Read More ›
projects – Bioinformatics
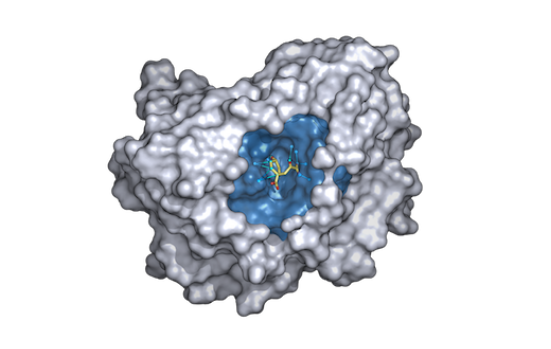 With the rise of computational modelling in the biotech industry, an emphasis is being put on the development of open-source toolkits for the simulation of drug-like molecular compounds in complex with proteins, fueling a rapid increase in computationally aided drug design. However, these workflows are often computationally expensive and are being migrated to the cloud. So what does cloud based drug discovery look like?
Read More ›
With the rise of computational modelling in the biotech industry, an emphasis is being put on the development of open-source toolkits for the simulation of drug-like molecular compounds in complex with proteins, fueling a rapid increase in computationally aided drug design. However, these workflows are often computationally expensive and are being migrated to the cloud. So what does cloud based drug discovery look like?
Read More ›
blog – Computational Chemistry
 Quantum mechanical (QM) methods provide a rigorous, physics-based framework for understanding molecular structure, properties, and reactivity at the atomic level.
Read More ›
Quantum mechanical (QM) methods provide a rigorous, physics-based framework for understanding molecular structure, properties, and reactivity at the atomic level.
Read More ›
projects – Data Science
 Can machine learning determine whether a wine is red or white based on its chemical composition? Can it predict whether a wine is good or bad? Using a dataset of 6,497 wines from the UCI Machine Learning Repository, this project explored wine classification and quality prediction. Binary Logistic Regression acheived a 97.3% accuracy in distinguishing between red and white wines. Further investigation using bayesian inference revealed optimal citric acid levels of 0.29 and 0.34 g.dm⁻³.
Read More ›
Can machine learning determine whether a wine is red or white based on its chemical composition? Can it predict whether a wine is good or bad? Using a dataset of 6,497 wines from the UCI Machine Learning Repository, this project explored wine classification and quality prediction. Binary Logistic Regression acheived a 97.3% accuracy in distinguishing between red and white wines. Further investigation using bayesian inference revealed optimal citric acid levels of 0.29 and 0.34 g.dm⁻³.
Read More ›
blog – Photochemistry
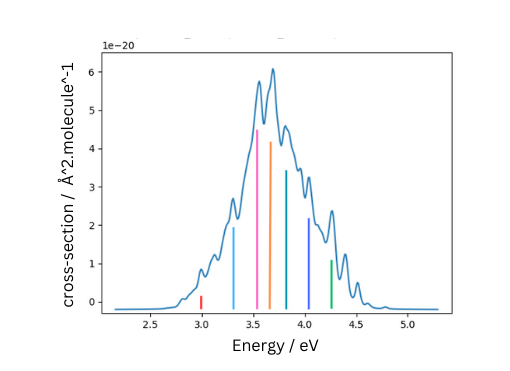 The Nuclear Ensemble Approach (NEA) is a computational chemistry approach used in photochemistry to model photoabsorption spectra and simulate nonadiabatic dynamics. By generating a representative set of molecular geometries (ensemble) from a Boltzmann-weighted distribution, NEM captures the effects of vibrational broadening and conformational flexibility on electronic transitions. Recent advancements, such as simulated annealing and optimal sampling, improve efficiency by reducing redundant configurations, enhancing accuracy in predicting UV-Vis absorption spectra and excited-state properties of gas-phase molecules and biomolecules.
Read More ›
The Nuclear Ensemble Approach (NEA) is a computational chemistry approach used in photochemistry to model photoabsorption spectra and simulate nonadiabatic dynamics. By generating a representative set of molecular geometries (ensemble) from a Boltzmann-weighted distribution, NEM captures the effects of vibrational broadening and conformational flexibility on electronic transitions. Recent advancements, such as simulated annealing and optimal sampling, improve efficiency by reducing redundant configurations, enhancing accuracy in predicting UV-Vis absorption spectra and excited-state properties of gas-phase molecules and biomolecules.
Read More ›
blog – Atmospheric Chemistry
 AtmoSpec is an innovative UV/Vis spectroscopy tool designed to streamline the calculation of photoabsorption cross-sections for volatile organic compounds (VOCs). Developed by the InSilico Photochemistry group at the University of Bristol, AtmoSpec enables users to input a molecule's SMILES code, select a theoretical framework, and perform quantum chemistry calculations using ORCA.
Read More ›
AtmoSpec is an innovative UV/Vis spectroscopy tool designed to streamline the calculation of photoabsorption cross-sections for volatile organic compounds (VOCs). Developed by the InSilico Photochemistry group at the University of Bristol, AtmoSpec enables users to input a molecule's SMILES code, select a theoretical framework, and perform quantum chemistry calculations using ORCA.
Read More ›
projects – Web development
 This website is an ongoing project, developing my abities in HTML and CSS. It aims to showcase my work, and help me track my development as a programmer and a chemist.
Read More ›
This website is an ongoing project, developing my abities in HTML and CSS. It aims to showcase my work, and help me track my development as a programmer and a chemist.
Read More ›
projects – Data Science
 The Nuclear Ensemble Method allows for exploring UV vis spectroscopy properties of small molecules accurately, however, it suffers from high computational burden due to large ensemble sizes, which become increasingly limiting with larger, more complex organic compounds. As such, methods for reducing intensity of calculations have been proposed. The Representative sampling method, aiming to reduce the subset while maintaining a high quality spectrum, was implemented as an automated workflow using Python.
Read More ›
The Nuclear Ensemble Method allows for exploring UV vis spectroscopy properties of small molecules accurately, however, it suffers from high computational burden due to large ensemble sizes, which become increasingly limiting with larger, more complex organic compounds. As such, methods for reducing intensity of calculations have been proposed. The Representative sampling method, aiming to reduce the subset while maintaining a high quality spectrum, was implemented as an automated workflow using Python.
Read More ›
projects – Cloud paradigms
 Scaling an ATLAS data processing script, going from a monolithic script to a containerized, service-based architecture. Functionality demonstrated locally using Docker swarm, with dynamic scaling of containers for fetching and processing tasks.
Read More ›
Scaling an ATLAS data processing script, going from a monolithic script to a containerized, service-based architecture. Functionality demonstrated locally using Docker swarm, with dynamic scaling of containers for fetching and processing tasks.
Read More ›
projects – Data Science
 How do you dunk your biscuit? Everyone seems to have an opinion, sometimes a strong one. Well it turns out that science does too, the Washburn equation for capillary action, has been shown to have great applicability to biscuits! This analysis presents a comprehensive description of biscuit dunking data to distinguish and predict brand of biscuit, as well as soaking time using physical data.
Read More ›
How do you dunk your biscuit? Everyone seems to have an opinion, sometimes a strong one. Well it turns out that science does too, the Washburn equation for capillary action, has been shown to have great applicability to biscuits! This analysis presents a comprehensive description of biscuit dunking data to distinguish and predict brand of biscuit, as well as soaking time using physical data.
Read More ›
projects – Data Visualization
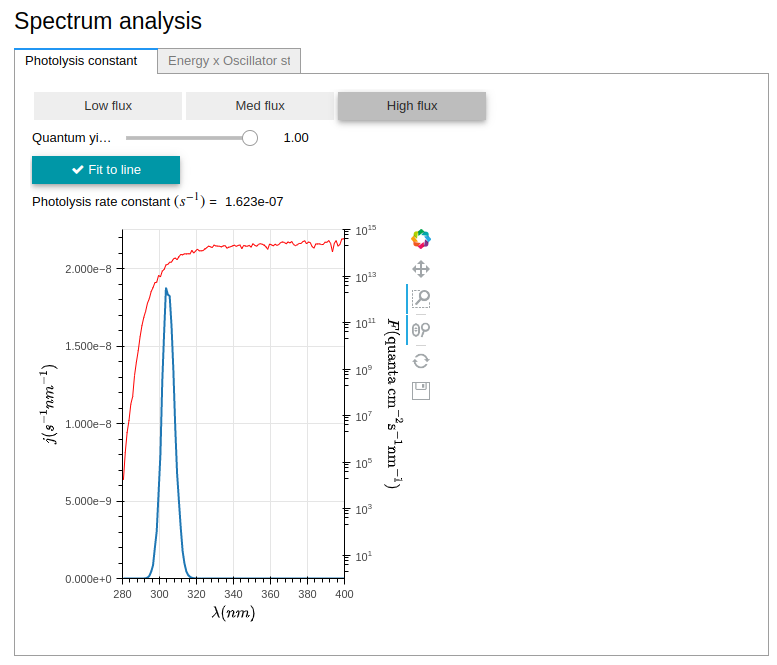 An interactive data visualization module, allowing for the exploration of photolysis properties in Atmospec. The photolysis rate constant was calculated using dynamic inputs from the user, and the graph is updated in real-time, allowing for researchers to interact with their results, and explore changes in data.
Read More ›
An interactive data visualization module, allowing for the exploration of photolysis properties in Atmospec. The photolysis rate constant was calculated using dynamic inputs from the user, and the graph is updated in real-time, allowing for researchers to interact with their results, and explore changes in data.
Read More ›
projects – Increasing Efficiency
 Exploring tools such as OpenMP, MPI and C++, as well as Python modules Cython and Numba to increase the efficiency of the computationally intensive Lebwhol-Lasher fluid model coded in Python
Read More ›
Exploring tools such as OpenMP, MPI and C++, as well as Python modules Cython and Numba to increase the efficiency of the computationally intensive Lebwhol-Lasher fluid model coded in Python
Read More ›
blog – Computational Chemistry
 Save the tedious lab work!
Read More ›
Save the tedious lab work!
Read More ›